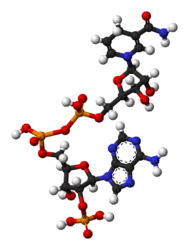
Nicotinamide adenine dinucleotide phosphate

| |
| Identifiers | |
|---|---|
3D model (JSmol)
|
|
| ChEBI | |
| ChEMBL | |
| ChemSpider | |
| ECHA InfoCard | 100.000.163 |
| MeSH | NADP |
PubChem CID
|
|
| UNII | |
| |
| |
| Properties | |
| C21H29N7O17P3 | |
| Molar mass | 744.416 g·mol−1 |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
| |
Nicotinamide adenine dinucleotide phosphate, abbreviated NADP[1][2] or, in older notation, TPN (triphosphopyridine nucleotide), is a cofactor used in anabolic reactions, such as the Calvin cycle and lipid and nucleic acid syntheses, which require NADPH as a reducing agent ('hydrogen source'). NADPH is the reduced form, whereas NADP+ is the oxidized form. NADP+ is used by all forms of cellular life. NADP+ is essential for life because it is needed for cellular respiration.[3]
NADP+ differs from NAD+ by the presence of an additional phosphate group on the 2' position of the ribose ring that carries the adenine moiety. This extra phosphate is added by NAD+ kinase and removed by NADP+ phosphatase.[4]
Biosynthesis
[edit]NADP+
[edit]In general, NADP+ is synthesized before NADPH is. Such a reaction usually starts with NAD+ from either the de-novo or the salvage pathway, with NAD+ kinase adding the extra phosphate group. ADP-ribosyl cyclase allows for synthesis from nicotinamide in the salvage pathway, and NADP+ phosphatase can convert NADPH back to NADH to maintain a balance.[3] Some forms of the NAD+ kinase, notably the one in mitochondria, can also accept NADH to turn it directly into NADPH.[5][6] The prokaryotic pathway is less well understood, but with all the similar proteins the process should work in a similar way.[3]
NADPH
[edit]NADPH is produced from NADP+. The major source of NADPH in animals and other non-photosynthetic organisms is the pentose phosphate pathway, by glucose-6-phosphate dehydrogenase (G6PDH) in the first step. The pentose phosphate pathway also produces pentose, another important part of NAD(P)H, from glucose. Some bacteria also use G6PDH for the Entner–Doudoroff pathway, but NADPH production remains the same.[3]
Ferredoxin–NADP+ reductase, present in all domains of life, is a major source of NADPH in photosynthetic organisms including plants and cyanobacteria. It appears in the last step of the electron chain of the light reactions of photosynthesis. It is used as reducing power for the biosynthetic reactions in the Calvin cycle to assimilate carbon dioxide and help turn the carbon dioxide into glucose. It has functions in accepting electrons in other non-photosynthetic pathways as well: it is needed in the reduction of nitrate into ammonia for plant assimilation in nitrogen cycle and in the production of oils.[3]
There are several other lesser-known mechanisms of generating NADPH, all of which depend on the presence of mitochondria in eukaryotes. The key enzymes in these carbon-metabolism-related processes are NADP-linked isoforms of malic enzyme, isocitrate dehydrogenase (IDH), and glutamate dehydrogenase. In these reactions, NADP+ acts like NAD+ in other enzymes as an oxidizing agent.[7] The isocitrate dehydrogenase mechanism appears to be the major source of NADPH in fat and possibly also liver cells.[8] These processes are also found in bacteria. Bacteria can also use a NADP-dependent glyceraldehyde 3-phosphate dehydrogenase for the same purpose. Like the pentose phosphate pathway, these pathways are related to parts of glycolysis.[3] Another carbon metabolism-related pathway involved in the generation of NADPH is the mitochondrial folate cycle, which uses principally serine as a source of one-carbon units to sustain nucleotide synthesis and redox homeostasis in mitochondria. Mitochondrial folate cycle has been recently suggested as the principal contributor to NADPH generation in mitochondria of cancer cells.[9]
NADPH can also be generated through pathways unrelated to carbon metabolism. The ferredoxin reductase is such an example. Nicotinamide nucleotide transhydrogenase transfers the hydrogen between NAD(P)H and NAD(P)+, and is found in eukaryotic mitochondria and many bacteria. There are versions that depend on a proton gradient to work and ones that do not. Some anaerobic organisms use NADP+-linked hydrogenase, ripping a hydride from hydrogen gas to produce a proton and NADPH.[3]
Like NADH, NADPH is fluorescent. NADPH in aqueous solution excited at the nicotinamide absorbance of ~335 nm (near UV) has a fluorescence emission which peaks at 445-460 nm (violet to blue). NADP+ has no appreciable fluorescence.[10]
Function
[edit]NADPH provides the reducing agents, usually hydrogen atoms, for biosynthetic reactions and the oxidation-reduction involved in protecting against the toxicity of reactive oxygen species (ROS), allowing the regeneration of glutathione (GSH).[11] NADPH is also used for anabolic pathways, such as cholesterol synthesis, steroid synthesis,[12] ascorbic acid synthesis,[12] xylitol synthesis,[12] cytosolic fatty acid synthesis[12] and microsomal fatty acid chain elongation.
The NADPH system is also responsible for generating free radicals in immune cells by NADPH oxidase. These radicals are used to destroy pathogens in a process termed the respiratory burst.[13] It is the source of reducing equivalents for cytochrome P450 hydroxylation of aromatic compounds, steroids, alcohols, and drugs.
Stability
[edit]NADH and NADPH are very stable in basic solutions, but NAD+ and NADP+ are degraded in basic solutions into a fluorescent product that can be used conveniently for quantitation. Conversely, NADPH and NADH are degraded by acidic solutions while NAD+/NADP+ are fairly stable to acid.[14][15]
Enzymes that use NADP(H) as a coenzyme
[edit]Many enzymes that bind NADP share a common super-secondary structure named named the "Rossmann fold". The initial beta-alpha-beta (βαβ) fold is the most conserved segment of the Rossmann folds. This segment is in contact with the ADP portion of NADP. Therefore, it is also called an "ADP-binding βαβ fold".[16]
- Adrenodoxin reductase: This enzyme is present ubiquitously in most organisms.[17] It transfers two electrons from NADPH to FAD. In vertebrates, it serves as the first enzyme in the chain of mitochondrial P450 systems that synthesize steroid hormones.[18]
Enzymes that use NADP(H) as a substrate
[edit]In 2018 and 2019, the first two reports of enzymes that catalyze the removal of the 2' phosphate of NADP(H) in eukaryotes emerged. First the cytoplasmic protein MESH1 (Q8N4P3),[19] then the mitochondrial protein nocturnin[20] were reported. Of note, the structures and NADPH binding of MESH1 (5VXA) and nocturnin (6NF0) are not related.
References
[edit]- ^ "NADP nicotinamide-adenine-dinucleotide phosphate". PubChem. U.S. National Library of Medicine. Retrieved 2024-08-22.
- ^ Karlson P (2014-05-12). Introduction to Modern Biochemistry. Academic Press. ISBN 978-1-4832-6778-4.
- ^ a b c d e f g Spaans SK, Weusthuis RA, van der Oost J, Kengen SW (2015). "NADPH-generating systems in bacteria and archaea". Frontiers in Microbiology. 6: 742. doi:10.3389/fmicb.2015.00742. PMC 4518329. PMID 26284036.
- ^ Kawai S, Murata K (April 2008). "Structure and function of NAD kinase and NADP phosphatase: key enzymes that regulate the intracellular balance of NAD(H) and NADP(H)". Bioscience, Biotechnology, and Biochemistry. 72 (4): 919–930. doi:10.1271/bbb.70738. PMID 18391451.
- ^ Iwahashi Y, Hitoshio A, Tajima N, Nakamura T (April 1989). "Characterization of NADH kinase from Saccharomyces cerevisiae". Journal of Biochemistry. 105 (4): 588–593. doi:10.1093/oxfordjournals.jbchem.a122709. PMID 2547755.
- ^ Iwahashi Y, Nakamura T (June 1989). "Localization of the NADH kinase in the inner membrane of yeast mitochondria". Journal of Biochemistry. 105 (6): 916–921. doi:10.1093/oxfordjournals.jbchem.a122779. PMID 2549021.
- ^ Hanukoglu I, Rapoport R (Feb–May 1995). "Routes and regulation of NADPH production in steroidogenic mitochondria". Endocrine Research. 21 (1–2): 231–241. doi:10.3109/07435809509030439. PMID 7588385.
- ^ Palmer M. "10.4.3 Supply of NADPH for fatty acid synthesis". Metabolism Course Notes. Archived from the original on 6 June 2013. Retrieved 6 April 2012.
- ^ Ciccarese F, Ciminale V (June 2017). "Escaping Death: Mitochondrial Redox Homeostasis in Cancer Cells". Frontiers in Oncology. 7: 117. doi:10.3389/fonc.2017.00117. PMC 5465272. PMID 28649560.
- ^ Blacker TS, Mann ZF, Gale JE, Ziegler M, Bain AJ, Szabadkai G, et al. (May 2014). "Separating NADH and NADPH fluorescence in live cells and tissues using FLIM". Nature Communications. 5 (1). Springer Science and Business Media LLC: 3936. Bibcode:2014NatCo...5.3936B. doi:10.1038/ncomms4936. PMC 4046109. PMID 24874098.
- ^ Rush GF, Gorski JR, Ripple MG, Sowinski J, Bugelski P, Hewitt WR (May 1985). "Organic hydroperoxide-induced lipid peroxidation and cell death in isolated hepatocytes". Toxicology and Applied Pharmacology. 78 (3): 473–483. Bibcode:1985ToxAP..78..473R. doi:10.1016/0041-008X(85)90255-8. PMID 4049396.
- ^ a b c d Rodwell V (2015). Harper's illustrated Biochemistry, 30th edition. USA: McGraw Hill. pp. 123–124, 166, 200–201. ISBN 978-0-07-182537-5.
- ^ Ogawa K, Suzuki K, Okutsu M, Yamazaki K, Shinkai S (October 2008). "The association of elevated reactive oxygen species levels from neutrophils with low-grade inflammation in the elderly". Immunity & Ageing. 5: 13. doi:10.1186/1742-4933-5-13. PMC 2582223. PMID 18950479.
- ^ Passonneau J (1993). Enzymatic analysis : a practical guide. Totowa, NJ: Humana Press. p. 3,10. ISBN 978-0-89603-238-5. OCLC 26397387.
- ^ Lu W, Wang L, Chen L, Hui S, Rabinowitz JD (January 2018). "Extraction and Quantitation of Nicotinamide Adenine Dinucleotide Redox Cofactors". Antioxidants & Redox Signaling. 28 (3): 167–179. doi:10.1089/ars.2017.7014. PMC 5737638. PMID 28497978.
- ^ Hanukoglu I (2015). "Proteopedia: Rossmann fold: A beta-alpha-beta fold at dinucleotide binding sites". Biochem Mol Biol Educ. 43 (3): 206–9. doi:10.1002/bmb.20849. PMID 25704928.
- ^ Hanukoglu I (December 2017). "Conservation of the Enzyme-Coenzyme Interfaces in FAD and NADP Binding Adrenodoxin Reductase-A Ubiquitous Enzyme". Journal of Molecular Evolution. 85 (5–6): 205–218. Bibcode:2017JMolE..85..205H. doi:10.1007/s00239-017-9821-9. PMID 29177972. S2CID 7120148.
- ^ Hanukoglu I (December 1992). "Steroidogenic enzymes: structure, function, and role in regulation of steroid hormone biosynthesis". The Journal of Steroid Biochemistry and Molecular Biology. 43 (8): 779–804. doi:10.1016/0960-0760(92)90307-5. PMID 22217824. S2CID 112729.
- ^ Ding CK, Rose J, Wu J, Sun T, Chen KY, Chen PH, et al. (2020). "MESH1 is a cytosolic NADPH phosphatase that regulates ferroptosis". Nature Metabolism. 2 (3): 270–277. bioRxiv 10.1101/325266. doi:10.1038/s42255-020-0181-1. PMC 7252213. PMID 32462112.
- ^ Estrella MA, Du J, Chen L, Rath S, Prangley E, Chitrakar A, et al. (2019). "The metabolites NADP+ and NADPH are the targets of the circadian protein Nocturnin (Curled)". Nature Communications. 10 (1): 2367. Bibcode:2019NatCo..10.2367E. bioRxiv 10.1101/534560. doi:10.1038/s41467-019-10125-z. PMC 6542800. PMID 31147539.